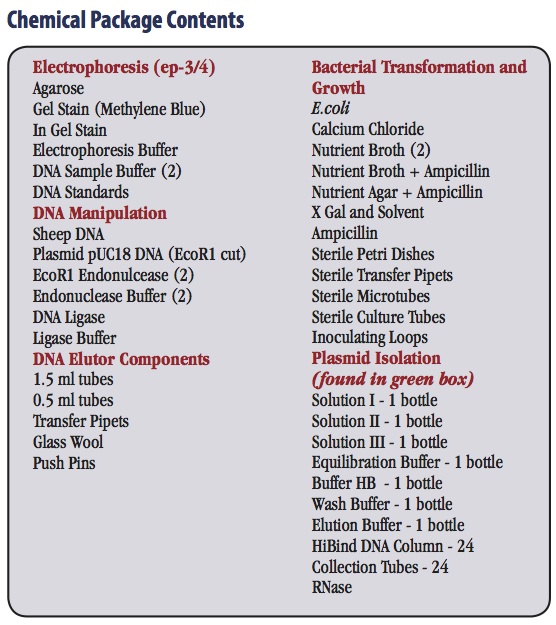
Description
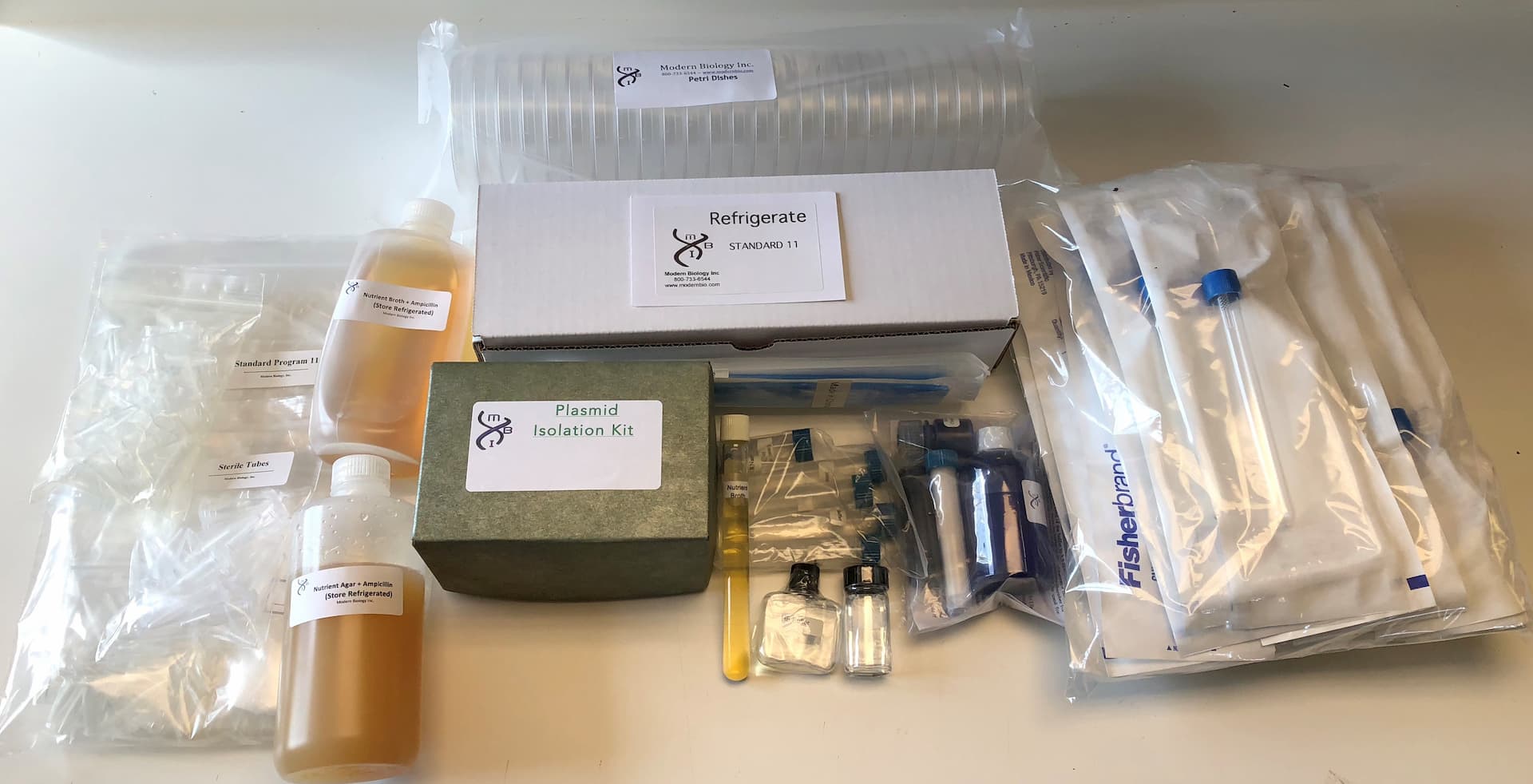
The sheep genome contains a highly repeated satellite sequence. In this program, students use recombinant DNA techniques to clone this satellite DNA and then characterize the sequence by restriction enzyme mapping anaylsis. The program provides essentially all of the instructions, chemicals, sterile media, and expendable accessories that are needed to carry out DNA electrophoresis, restriction nuclease digestion, DNA ligations, bacterial transformations, bacterial selection for ampicillin resistance and ß – glactosidase production and plasmid isolation. A microcentrifuge is required for the program.
Sample
Lab Session 1
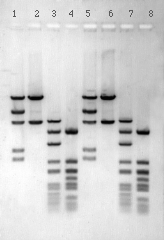
Students digest sheep DNA with EcoR1 and electrophorese the DNA fragments on an agarose gel in order to seperate the satellite DNA from the remaining sequences in the sheep genome.
Lab Sessions 2-3
The satellite DNA band is eluted from the gel and ligated to EcoR1-digested pUC18 in order to create a recombinant plasmid that contains the satellite sequence. The recombinant plasmid is then introduced into competent E.coli cells by transformation and colonies containing foreign DNA are identified by a blue-white color selection assay.
Lab Sessions 4-5
Students prepare overnight cultures from white colonies which should contain the sheep satellite sequence. Plasmids are then isolated from the cultures by a mini-prep procedure that yields high quality plasmid DNA without the use of toxic organic solvents. Students digest the plasmids with EcoR1 and determine the size of the DNA fragments by electrophoresis. The results of this gel analysis enable the student to identify the satellite sequence in the recombinant plasmid.
 Due to Customs restrictions, we only accept orders from educational institutions within the Continental United States, Alaska or Hawaii.
Due to Customs restrictions, we only accept orders from educational institutions within the Continental United States, Alaska or Hawaii.